TOP 10 JUTSUS OF FEATURE ENGINEERING EVERY DATA SCIENTISTS NINJA SHOULD KNOW RIGHT NOW
If you want to be a successful data scientist and to make your model predict the most accurate result, then the following article is for you.
What is Feature Engineering?
As per Wikipedia, Feature engineering is the process of using domain
knowledge to extract features from raw data via data
mining techniques. These features can be used to improve
the performance of machine learning algorithms. Feature engineering can be considered as applied
machine learning itself.
Okay, so we
saw what Wikipedia said about it but still, we want to know more about it. So
let’s divide this term into 2 parts for better understanding i.e Feature and
Engineering.
What are the
Features in a dataset?
Basically,
all machine learning algorithms use some input data (independent data) to
generate output. These input data are features, which are usually in the form of
structured columns.
So, why do we engineer it?
To get the most accurate or I can say to get the most precise output from our machine learning the algorithm we need to give it cleanest data possible which should be compatible with our machine learning algorithm.
So the process of preparing the proper input dataset for our machine learning model is known as Feature Engineering.
Link of the complete survey by Forbes.
Data scientists spend 60% of their time cleaning and organizing data. Collecting data sets come second at 19% of their time, meaning data scientists spend around 80% of their time on preparing and managing data for analysis.
76% of data
scientists view data preparation as the least enjoyable part of their work
57% of data scientists regard cleaning and organizing data as the least enjoyable part of their work and 19% say this about collecting data sets.
List of
Techniques you can find in this blog:
1.) Techniques
of Imputation of numerical and categorical data
2.) Dealing
with outliers
3.) Binning
4.) Techniques
of dealing with Gaussian-Distribution / Skewness
5.) OneHotEncoding
and OrdinalEncoder
6.) Feature
splitting & extraction
7.) Group By
8.) Concat,
Merge, Join
9.) Scaling
10.) Extracting
Date
1.) Techniques of Imputation of numerical and categorical data
In this we deal with missing values that are present in our dataset.
The most simple solution to deal with missing values is to drop that row or column, suppose that you have more than 75% of values missing that you can definitely drop that row or column.
threshold = 0.75
# Columns:
data = data[data.columns[data.isnull().mean() < threshold]]
#------------------------------------------------------------------------
# Rows:
data = data.loc[data.columns[data.isnull().mean(axis=1) < threshold]]
# fill NaN values with 0
data = data.fillna(0)
#------------------------------------------------
# fill NaN values with median
data = data.fillna(data.mean())
#------------------------------------------------
# fill NaN values with median: Better than Mean
data = data.fillna(data.median())
#------------------------------------------------
# fill NaN values with mode
data = data.fillna(data.mode())
(i) Simple
Imputer
-A simple imputer is univariate which means it will take only a single feature in the count.
# using simple imputer
import numpy as np
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(missing_values=np.nan, strategy='median') # can be mean, median, mode
imputer = imputer.fit(x)
data = imputer.transform(x)
print('Imputed Data:',data)
-What if the
feature “x” which has NaN values is very well co-correlated with the features
such as “y” or “z”. Let’s say, people with higher “Age” give more “Rent” and
people with lower “Age” give less “Rent”. So that suggests a multivariate
approach. Which just means it takes multiple features into count.
-So the Iterative Imputer and KNN Imputer comes into the concentration.
(ii) Iterative
Imputer
-In
iterative imputer for all the rows in which “Age” is not missing sklearn trains
a regression model.
-Where other
features such as “Gender” and “Rent” are
considered as independent input features and “Age” is considered as the target
output feature.
-Now for all rows in which “Age” is missing it makes predictions for “Age” by using features “Gender” and “Rent” to trained model.
# IterativeImputer
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
imputer_it = IterativeImputer()
imputer_it.fit_transform(x)
NOTE: This model used by iterative imputer is totally independent of the model you are using this dataset as training data.
(iii) KNN
Imputer
-Let’s
pretend we have a row in which “Age” is missing.
-Then sklearn finds the 2 most similar rows measured by how close the “Gender” and “Fare” values are to this row.
# KNNImputer
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import KNNImputer
imputer_knn = KNNImputer(n_neighbors=2)
imputer_knn.fit_transform(x)
Dealing with the categorical and nominal data with the help of mode
data['categorical_col'] = data['categorical_col'].fillna(data['categorical_col'].mode()[0])
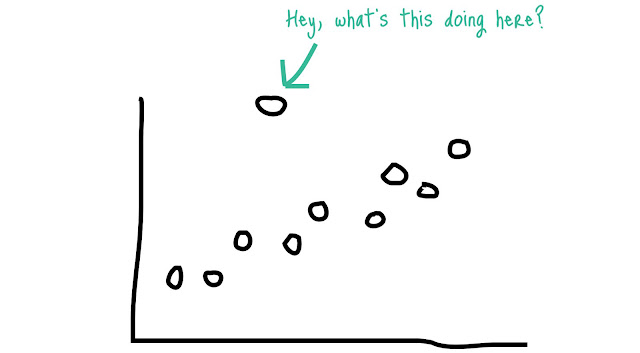
2.) Dealing with outliers
Detecting
outliers:
Using
Z-score:
Data that
fall outside of the 3rd standard deviation is considered as an outlier. We can use a Z-score if Z-score falls outside of μ+3 or μ-3 then we will consider it as an outlier.
 |
| Z-Score |
import numpy as np
box = [10,20,15,25,11,12,16,26,19,29,3000]
outliers = []
def detect_outliers(data):
Threshold = 3
mean = np.mean(data)
std = np.std(data)
for i in data:
z_score = (i-mean)/std
if np.abs(z_score) > Threshold:
outliers.append(i)
return outliers
out = detect_outliers(box)
out
Using IQR:
Datapoint
that fall outside of 1.5 times od Inter Quartile Range above 1st the quartile and the 3rd
quartile.
import numpy as np
box = [10,20,15,25,11,12,16,26,19,29,3000]
box = sorted(box)
Q1,Q3 = np.percentile(box, [25,75])
print(Q1,Q3)
iqr = Q3 - Q1
print(iqr)
# Lower bound and higher bound values
lower_bound_val = Q1 - (1.5*iqr)
higher_bound_val = Q3 + (1.5*iqr)
print(lower_bound_val,higher_bound_val)
Using Box-plot:
The values
which are beyond the Max or Min are considered as the outliers.
for feature in data:
dataset = data.copy()
if 0 in dataset[feature].unique():
pass
else:
dataset[feature] = np.log(dataset[feature])
dataset.boxplot(column=feature)
plt.ylabel(feature)
plt.title(feature)
plt.show()
fact = 3
upper_bound_val = data['column'].mean() + data['column'].std() * fact
lower_bound_val = data['column'].mean() - data['column'].std() * fact
data = data[(data['column'] < upper_bound_val) & (data['column'] > lower_bound_val)]
data = pd.DataFrame(np.random.randn(500,4))
from scipy import stats
data[(np.abs(stats.zscore(data)) < 3).all(axis=1)]
upper_bound_val = data['column'].quantile(0.95)
lower_bound_val = data['column'].quantile(0.05)
data = data[(data['column'] < upper_bound_val) & (data['column'] > lower_bound_val)]
3.) Binning
Image Credit: https://wisdomschema.com/data-binning/
-Binning can
be applied to both categorical and numerical data.
- The main reason behind binning is to make the model more robust and prevent overfitting,
however, it has a performance cost.
#Numerical
Binning Example
Value Bin
0-30 =>
Fail
31-70 =>
Average
71-100 => Excelent
#Categorical
Binning Example
Value Bin
Mumbai =>
Maharashtra
Pune =>
Maharashtra
Bikaner =>
Rajasthan
Jaipur =>
Rajasthan
Numerical
Binning Example:
data['bin'] = pd.cut(data['value'], bins=[0,30,70,100], labels=["Fail", "Average", "Excelent"])
value | bin
0 | 2 | Fail
1 | 45 | Average
2 | 7 | Fail
3 | 85 | Excellent
4 | 28 | Fail
Categorical
Binning Example:
conditions = [
data['State'].str.contains('Mumbai'),
data['State'].str.contains('Pune'),
data['State'].str.contains('Bikaner'),
data['State'].str.contains('Jaipur')]
choices = ['Maharashtra', 'Maharashtra', 'Rajasthan', 'Rajasthan']
data['Continent'] = np.select(conditions, choices, default='Other')
value | bin 0 | Mumbai | Maharashtra
1 | Pune | Maharashtra
2 | Bikaner | Rajasthan
3 | Delhi | Other
4 | Jaipur | Rajasthan
4.)Techniques of dealing with Gaussian-Distribution / Skewness
-Helps to
handles skewed data and after transformation, the distribution becomes more
approximate to the normal.
-Decreases
the effect of the outliers to the normalization of magnitude difference and the model becomes more robust.
(i)Log
Transformation:
Log transformation is a
data transformation method in which it replaces by log(x) with base
10, base 2, or natural log.
data['log_column'] = np.log(data['column']+1)
(ii)Reciprocal Transformation:
In Reciprocal Transformation, x
will replace by the inverse of X(1/X). The reciprocal transformation will give
little effect on the shape of the distribution. This transformation can be only
used for non-zero values. The skewness for the transformed data is increased.
data['Reciprocal_column'] = 1/(data['column']+1)
(iii)Square-Root Transformation:
This transformation will give a moderate effect on distribution. The main advantage of square root transformation is, it can be applied to
zero values. Here the x will replace by the square root(x).
It is weaker than the Log Transformation.
data['sqr_column'] = data['column']**(1/2)
5.)OneHotEncoding and OrdinalEncoder
Since our computer systems only understand numerical data. So categorical data makes no sense for them unless they are converted into the numerical data and for that we use techniques such as OneHotEncoding and OrdinalEncoders.
# For single
column
encoded_col = pd.get_dummies(data['column'], drop_first = True)
encoded_multi_col = pd.get_dummies(data,columns = ['column1', 'column2', 'columnN'], drop_first=True)
For unordered
(Nominal) data we will be using OneHotEncoding
Example: Male,
Female
# OneHotEncoding is for unordered (Nominal) data
# Example: Male, Female
from sklearn.preprocessing import OneHotEncoder
ohe = OneHotEncoder(sparse=False)
ohe.fit_transform(data[['column']])
For ordered
(Ordinal) data we will be using OrdinalEncoder
Example: First,
Second, Third
# OrdinalEncoder is for ordered (Ordinal) data
# Example: First, Second, Third
from sklearn.preprocessing import OrdinalEncoder
oe = OrdinalEncoder(categories=[['First', 'Second', 'Third'],
['O','A','B','C']])
oe.fit_transform(data[['Rank', 'Grade']])
6.)Feature splitting & extraction
-By extracting
the utilizable part of a column into a new feature:
*We enable the machine-learning algorithms to comprehend them.
*Make possible
to bin and group them.
*Improve model
performance by uncovering potential information.
Splitting a function is a good option, but however, there is one way of splitting features.
It depends on the characteristics of the column, how to split it.
Feature
Splitting:
import pandas as pd
import numpy as np
data = [('Arnold Schwarzenegger','M'),
('Natasha Romanova','F'),
('Sylvester Stallone','M'),
('Gal Gadot','F'),
('Dwayne Johnson','M')]
df = pd.DataFrame(data, columns=['name', 'gender'])
df
# Extracting First Name
df.name.str.split(" ").map(lambda X : X[0])
# Extracting Last Name
df.name.str.split(" ").map(lambda X : X[-1])
Feature Extraction:
weather_data = [('1/1/2017', 32, 6, 'Rain'),
('1/2/2017', 30, 7, 'Sunny'),
('1/3/2017', 32, 2, 'Snow'),
('1/4/2017', 34, 6, 'Snow'),
('1/5/2017', 32, 4, 'Rain'),
('1/6/2017', 32, 2, 'Sunny')
]
df = pd.DataFrame(weather_data, columns=['day', 'temp', 'windspeed', 'weather'])
df
df['day'][df['weather']=='Rain']
df.day[df.temp == df.temp.max()]
7.) Group By
groupby() function is used to split the data into groups based on some criteria. pandas objects can be split on any of their axes. The abstract definition of grouping is to provide a mapping of labels to group names.
weather_data = [('1/1/2017', 'Mumbai' , 32, 6, 'Rain'),
('1/2/2017', 'Pune' , 30, 7, 'Sunny'),
('1/3/2017', 'Mumbai' , 32, 2, 'Snow'),
('1/4/2017', 'Pune' , 34, 6, 'Snow'),
('1/5/2017', 'Mumbai' , 32, 4, 'Rain'),
('1/6/2017', 'Delhi' , 32, 2, 'Sunny')
]
data = pd.DataFrame(weather_data, columns=['day', 'city', 'temp', 'windspeed', 'weather'])
data
grp_city = data.groupby('city')
for city, city_data in grp_city:
print(city)
print(city_data)
# Get specific group
grp_city.get_group('Mumbai')
print(grp_city.max())
print(grp_city.mean())
print(grp_city.describe())
8.) Concat, Merge, Join
Works like how
we had worked on SQL.
Maharashtra_weather_data = pd.DataFrame({
'city': ['Mumbai', 'Pune', 'Thane'],
'temperature': [32, 45, 30],
'humidity': [80, 60, 78]
})
Maharashtra_weather_data
Gujarat_weather_data = pd.DataFrame({
'city': ['Surat', 'Rajkot', 'Mehsana'],
'temperature': [21, 24, 35],
'humidity': [68, 65, 75]
})
Gujarat_weather_data
(i) Concat:
-Column Wise
Concatenation
df = pd.concat([Maharashtra_weather_data, Gujarat_weather_data])
df
-Row Wise Concatenation
df = pd.concat([Maharashtra_weather_data, Gujarat_weather_data], axis=1)
df
(ii) Merge
temp_data = pd.DataFrame({
'city':['Mumbai','Delhi','Banglore','Hydrabad'],
'temp':[32,45,30,40]
})
temp_data
humidity_data = pd.DataFrame({
'city':['Mumbai','Delhi','Banglore'],
'humidity':[62,65,70]
})
humidity_data
# Merge 2 dataframe without explicitly mentioning the index
df = pd.merge(temp_data, humidity_data, on='city')
df
(iii) Join
df = pd.merge(temp_data, humidity_data, on='city',how='outer')
df
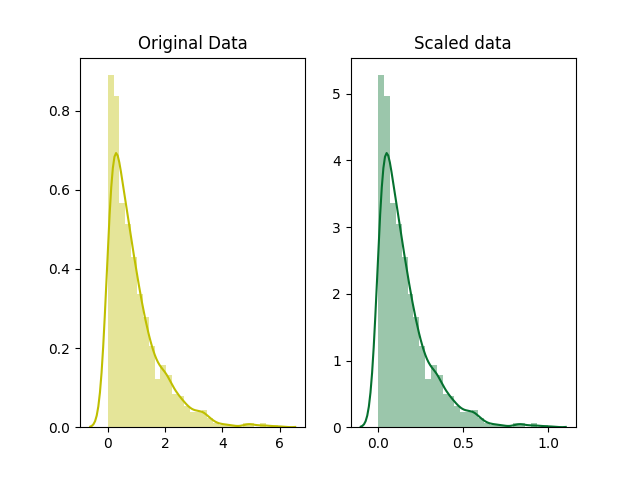
9.) Scaling
Scaling are done basically in 2 ways first is
Normalization and the second is Standardization.
data = pd.DataFrame({
'name':['Ram', 'Lakhan', 'Shiva', 'Ria', 'Lucy', 'Suraj', 'Rohan', 'Anny', 'Priya', 'Niraj'],
'age':[25,40,33,26,30,35,28,43,36,50],
'Salary':[25000, 32000, 50000, 33000, 20000, 29000, 22000, 52000, 23000, 26000],
'purchased':[1,0,1,1,0,0,0,1,0,1]
})
data
(i) Normalization
(MinMaxScaler)
* X_norm = X – X_min
/ X_max – X_min
* Range 0 to 1
* Due to the decrease in standard deviation the effect of outliers increases.
*Before
Normalization, it is recommended to handle the outliers.
from sklearn.model_selection import train_test_split
X = data.drop(['name','purchased'], axis=1)
y = data['purchased']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
from sklearn.preprocessing import MinMaxScaler
Min_Max_scaler = MinMaxScaler()
Min_Max_X_train = Min_Max_scaler.fit_transform(X_train)
Min_Max_X_test = Min_Max_scaler.transform(X_test)
Min_Max_X_train
Min_Max_X_test
(ii) Standardization
(StandardScaler)
*Z = X - μ / σ
*μ = 0 and σ = 1
*Range -1 to 1
*If the standard
deviation of feature is different, their range also would differ from each
other.
from sklearn.preprocessing import StandardScaler
StandardScaler_scaler = StandardScaler()
StandardScaler_X_train = StandardScaler_scaler.fit_transform(X_train)
StandardScaler_X_test = StandardScaler_scaler.transform(X_test)
StandardScaler_X_train
StandardScaler_X_test
10.) Extracting Date
There are 3 ways we can preprocessing date
*Extracting the parts of the date
into the different columns: YEAR, MONTH, DAY.
*Extracting the time period between
the current date and column in terms of YEAR, MONTH, DAY.
*Extracting some specific features
from the date such as the name of the weekday, weekend, etc.
*With dealing with date features
like that our machine learning model can easily understand data and deal with
the data.
from datetime import date
data = pd.DataFrame({
'date': ['01/01/2017', '04/12/2000', '23/04/2011', '11/02/2008', '08/08/2018']
})
data
data.info()
# Transform String to Date
data['date'] = pd.to_datetime(data.date, format='%d/%m/%Y')
data.info()
# Extracat year
data['year'] = data['date'].dt.year
data
# Extract month
data['month'] = data['date'].dt.month
data
# Extract day
data['day'] = data['date'].dt.day
data
# Extracting passed year since the date
data['passed_years'] = date.today().year - data['date'].dt.year
data
# Extracting passed month since the date
data['passed_months'] = (date.today().year - data['date'].dt.year)*12 + (date.today().month - data['date'].dt.month)
data
Here is the Github link of all the code which is mentioned here:
Github Link of the Notebook: Link
Use all the tips and tricks which are mentioned above and you will surely get amazing data to train your model on.
So we hope that you enjoyed this session. If you did then please share it with your friends and spread this knowledge.
Follow us at :
Instagram :
https://www.instagram.com/infinitycode_x/
Facebook :
https://www.facebook.com/InfinitycodeX/
Twitter :
https://twitter.com/InfinityCodeX1


















































Excellent article... Thank you for providing such valuable information; the contents are quite intriguing. I'll be waiting for the next post on Big Data Engineering Services with great excitement.
ReplyDelete